Molybdenum »
PDB 1aa6-1n61 »
1jro »
Molybdenum in PDB 1jro: Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus
Enzymatic activity of Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus
All present enzymatic activity of Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus:
1.1.1.204;
1.1.1.204;
Protein crystallography data
The structure of Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus, PDB code: 1jro
was solved by
J.J.Truglio,
K.Theis,
S.Leimkuhler,
R.Rappa,
K.V.Rajagopalan,
C.Kisker,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 50.00 / 2.70 |
| Space group | P 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 92.875, 141.053, 158.113, 109.53, 105.83, 101.33 |
| R / Rfree (%) | 21.3 / 25.2 |
Other elements in 1jro:
The structure of Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus also contains other interesting chemical elements:
| Iron | (Fe) | 16 atoms |
| Calcium | (Ca) | 4 atoms |
Molybdenum Binding Sites:
The binding sites of Molybdenum atom in the Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus
(pdb code 1jro). This binding sites where shown within
5.0 Angstroms radius around Molybdenum atom.
In total 4 binding sites of Molybdenum where determined in the Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus, PDB code: 1jro:
Jump to Molybdenum binding site number: 1; 2; 3; 4;
In total 4 binding sites of Molybdenum where determined in the Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus, PDB code: 1jro:
Jump to Molybdenum binding site number: 1; 2; 3; 4;
Molybdenum binding site 1 out of 4 in 1jro
Go back to
Molybdenum binding site 1 out
of 4 in the Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus
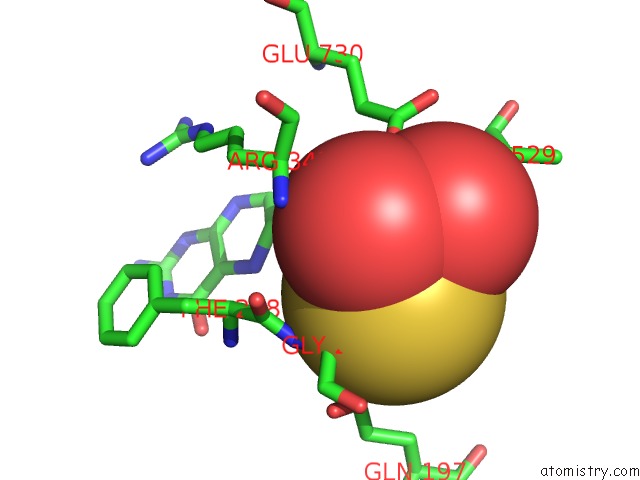
Mono view
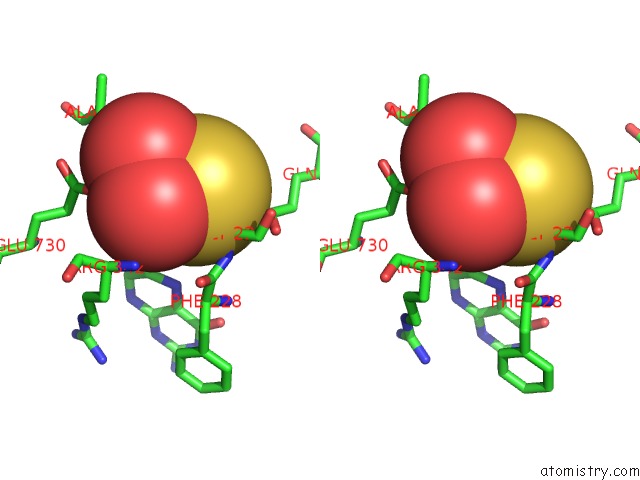
Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 1 of Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus within 5.0Å range:
|
Molybdenum binding site 2 out of 4 in 1jro
Go back to
Molybdenum binding site 2 out
of 4 in the Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus
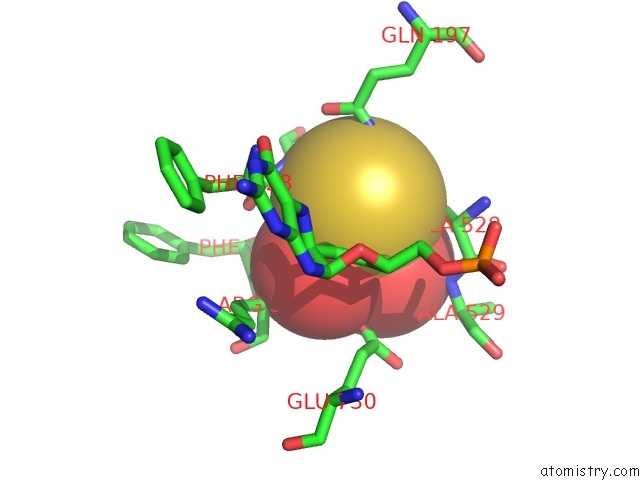
Mono view
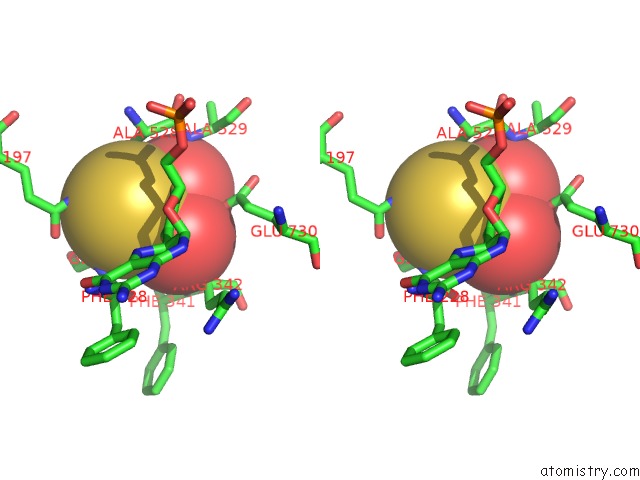
Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 2 of Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus within 5.0Å range:
|
Molybdenum binding site 3 out of 4 in 1jro
Go back to
Molybdenum binding site 3 out
of 4 in the Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus
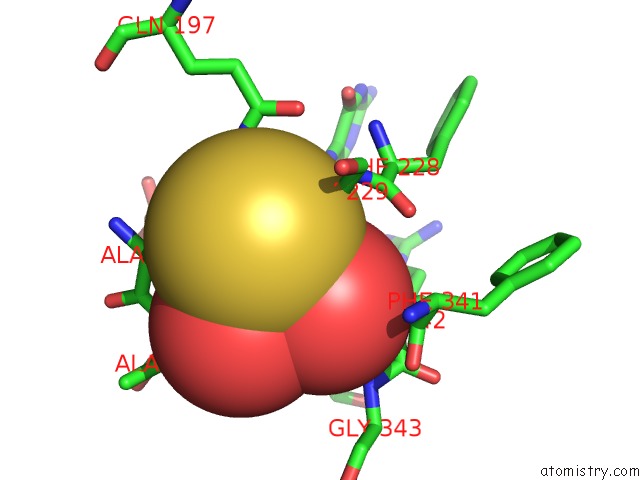
Mono view
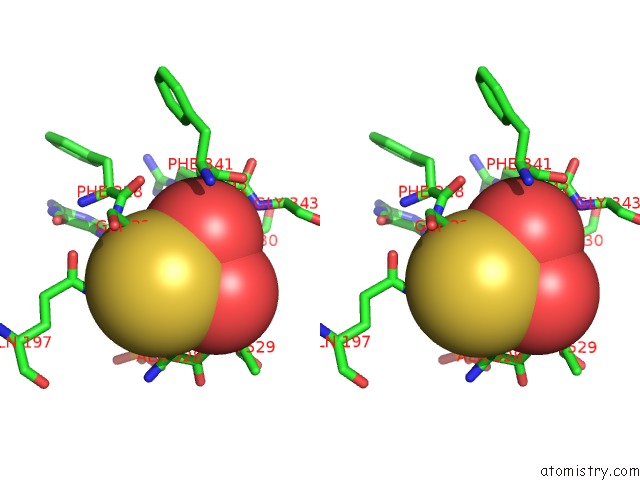
Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 3 of Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus within 5.0Å range:
|
Molybdenum binding site 4 out of 4 in 1jro
Go back to
Molybdenum binding site 4 out
of 4 in the Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus
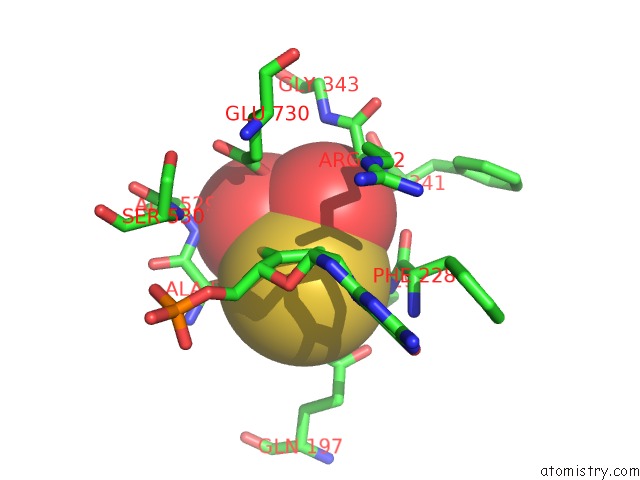
Mono view
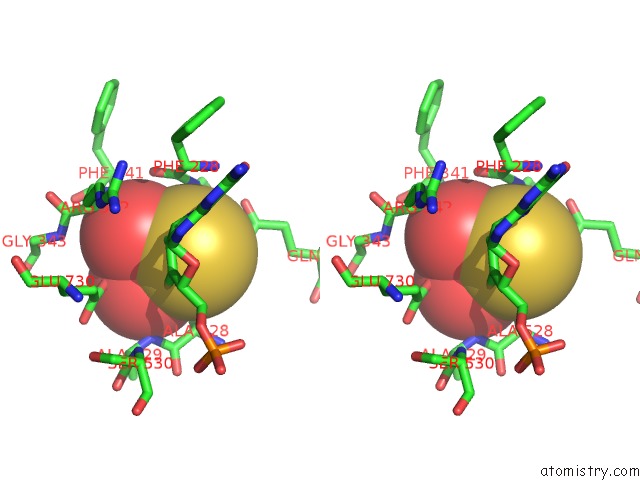
Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 4 of Crystal Structure of Xanthine Dehydrogenase From Rhodobacter Capsulatus within 5.0Å range:
|
Reference:
J.J.Truglio,
K.Theis,
S.Leimkuhler,
R.Rappa,
K.V.Rajagopalan,
C.Kisker.
Crystal Structures of the Active and Alloxanthine-Inhibited Forms of Xanthine Dehydrogenase From Rhodobacter Capsulatus Structure V. 10 115 2002.
ISSN: ISSN 0969-2126
PubMed: 11796116
DOI: 10.1016/S0969-2126(01)00697-9
Page generated: Sun Aug 17 02:55:16 2025
ISSN: ISSN 0969-2126
PubMed: 11796116
DOI: 10.1016/S0969-2126(01)00697-9
Last articles
Na in 3QFHNa in 3QG6
Na in 3QFD
Na in 3QF9
Na in 3QEM
Na in 3QEL
Na in 3QC5
Na in 3QB9
Na in 3QB5
Na in 3Q9F