Molybdenum »
PDB 1n62-2c9x »
1xdq »
Molybdenum in PDB 1xdq: Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase
Protein crystallography data
The structure of Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase, PDB code: 1xdq
was solved by
L.Loschi,
S.J.Brokx,
T.L.Hills,
G.Zhang,
M.G.Bertero,
A.L.Lovering,
J.H.Weiner,
N.C.Strynadka,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 39.75 / 2.55 |
| Space group | I 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 110.345, 165.064, 181.447, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 23 / 27.4 |
Molybdenum Binding Sites:
The binding sites of Molybdenum atom in the Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase
(pdb code 1xdq). This binding sites where shown within
5.0 Angstroms radius around Molybdenum atom.
In total 5 binding sites of Molybdenum where determined in the Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase, PDB code: 1xdq:
Jump to Molybdenum binding site number: 1; 2; 3; 4; 5;
In total 5 binding sites of Molybdenum where determined in the Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase, PDB code: 1xdq:
Jump to Molybdenum binding site number: 1; 2; 3; 4; 5;
Molybdenum binding site 1 out of 5 in 1xdq
Go back to
Molybdenum binding site 1 out
of 5 in the Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase
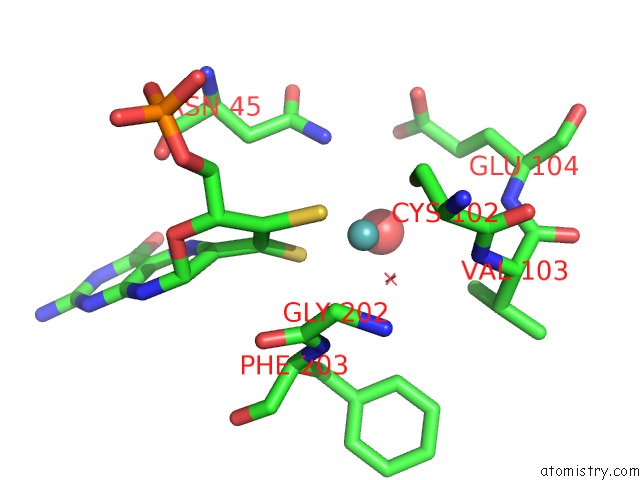
Mono view
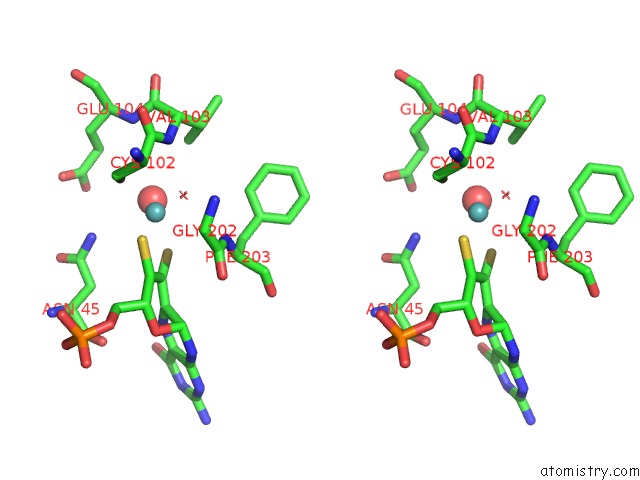
Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 1 of Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase within 5.0Å range:
|
Molybdenum binding site 2 out of 5 in 1xdq
Go back to
Molybdenum binding site 2 out
of 5 in the Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase
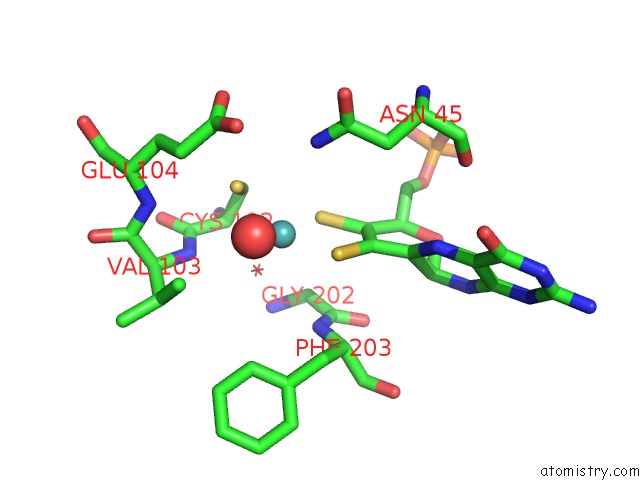
Mono view
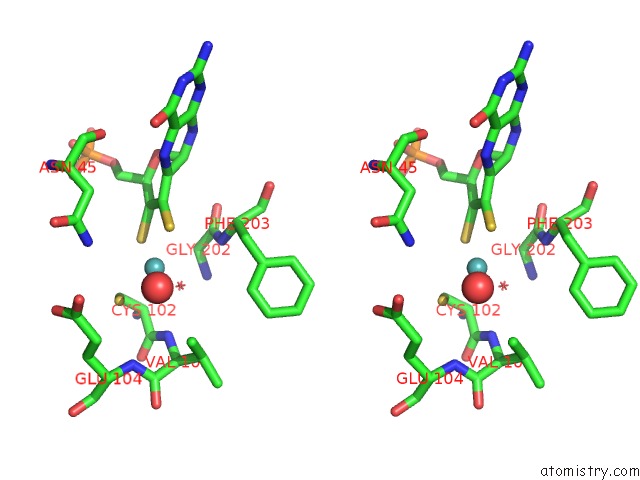
Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 2 of Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase within 5.0Å range:
|
Molybdenum binding site 3 out of 5 in 1xdq
Go back to
Molybdenum binding site 3 out
of 5 in the Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase
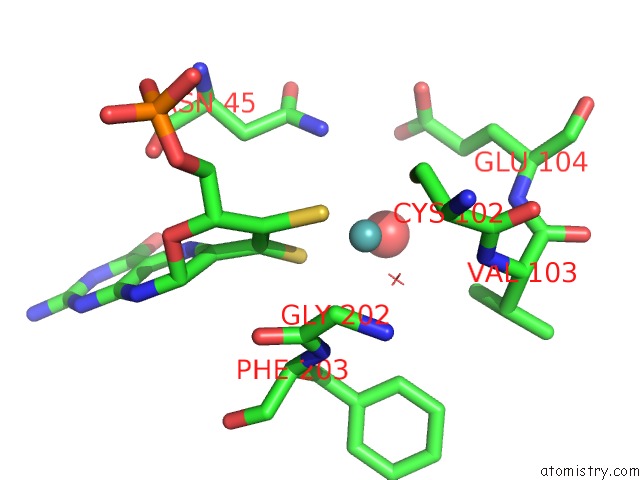
Mono view
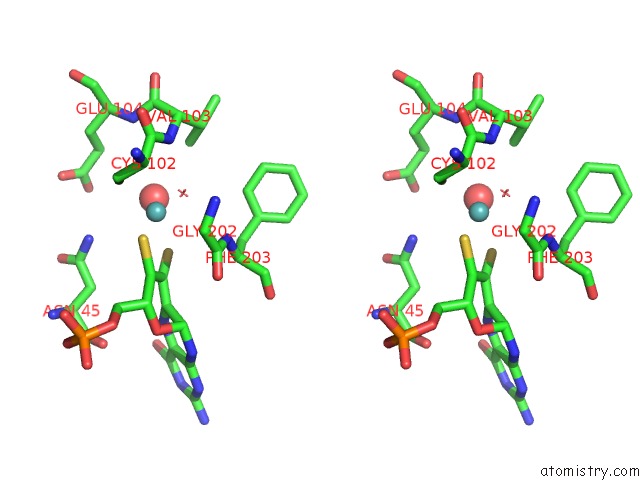
Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 3 of Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase within 5.0Å range:
|
Molybdenum binding site 4 out of 5 in 1xdq
Go back to
Molybdenum binding site 4 out
of 5 in the Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase
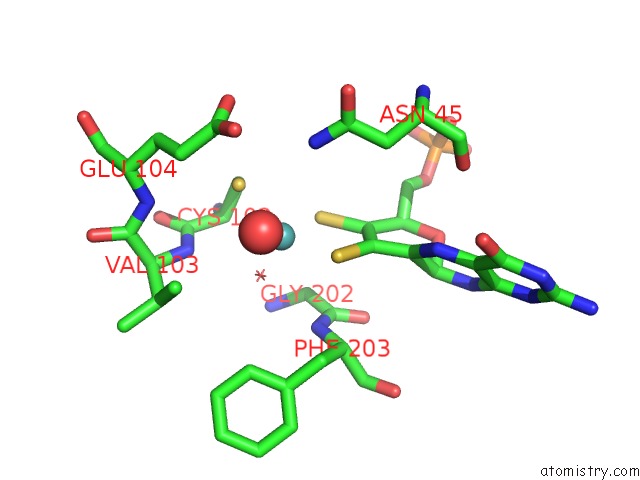
Mono view
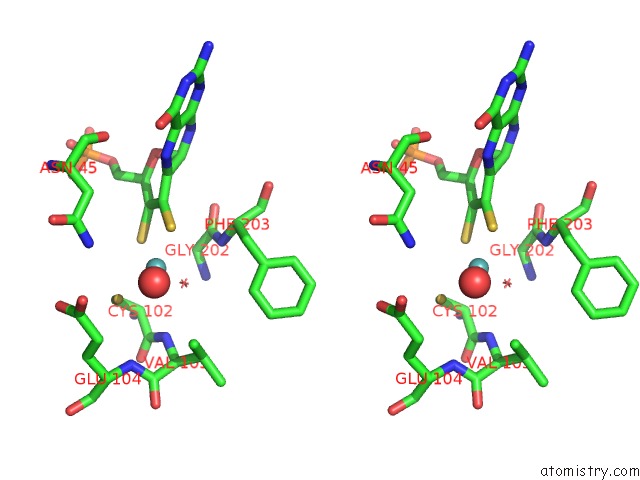
Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 4 of Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase within 5.0Å range:
|
Molybdenum binding site 5 out of 5 in 1xdq
Go back to
Molybdenum binding site 5 out
of 5 in the Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase
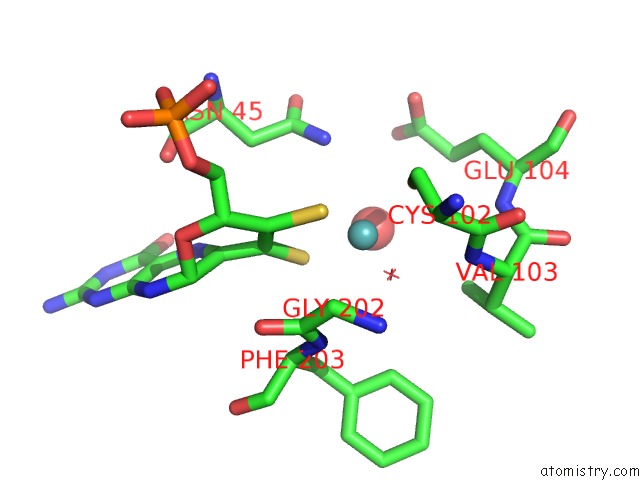
Mono view
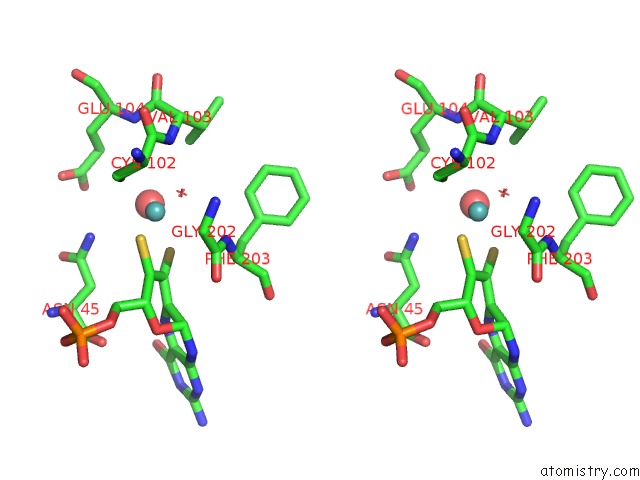
Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 5 of Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase within 5.0Å range:
|
Reference:
L.Loschi,
S.J.Brokx,
T.L.Hills,
G.Zhang,
M.G.Bertero,
A.L.Lovering,
J.H.Weiner,
N.C.Strynadka.
Structural and Biochemical Identification of A Novel Bacterial Oxidoreductase. J.Biol.Chem. V. 279 50391 2004.
ISSN: ISSN 0021-9258
PubMed: 15355966
DOI: 10.1074/JBC.M408876200
Page generated: Sun Aug 17 03:03:07 2025
ISSN: ISSN 0021-9258
PubMed: 15355966
DOI: 10.1074/JBC.M408876200
Last articles
Na in 5VY2Na in 5VYB
Na in 5VYL
Na in 5VU0
Na in 5VWN
Na in 5VXA
Na in 5VS2
Na in 5VRA
Na in 5VS3
Na in 5VRZ