Molybdenum »
PDB 1n62-2c9x »
2bii »
Molybdenum in PDB 2bii: Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta
Enzymatic activity of Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta
All present enzymatic activity of Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta:
1.7.1.2;
1.7.1.2;
Protein crystallography data
The structure of Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta, PDB code: 2bii
was solved by
K.Fischer,
G.Barbier,
H.-J.Hecht,
R.R.Mendel,
W.H.Campbell,
G.Schwarz,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 87.71 / 1.70 |
| Space group | C 2 2 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 122.722, 123.080, 149.509, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 18.5 / 21.2 |
Other elements in 2bii:
The structure of Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta also contains other interesting chemical elements:
| Sodium | (Na) | 2 atoms |
Molybdenum Binding Sites:
The binding sites of Molybdenum atom in the Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta
(pdb code 2bii). This binding sites where shown within
5.0 Angstroms radius around Molybdenum atom.
In total 2 binding sites of Molybdenum where determined in the Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta, PDB code: 2bii:
Jump to Molybdenum binding site number: 1; 2;
In total 2 binding sites of Molybdenum where determined in the Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta, PDB code: 2bii:
Jump to Molybdenum binding site number: 1; 2;
Molybdenum binding site 1 out of 2 in 2bii
Go back to
Molybdenum binding site 1 out
of 2 in the Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta
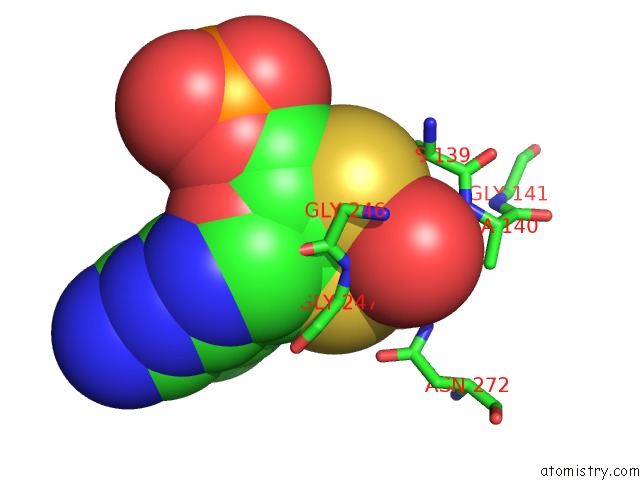
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 1 of Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta within 5.0Å range:
|
Molybdenum binding site 2 out of 2 in 2bii
Go back to
Molybdenum binding site 2 out
of 2 in the Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta
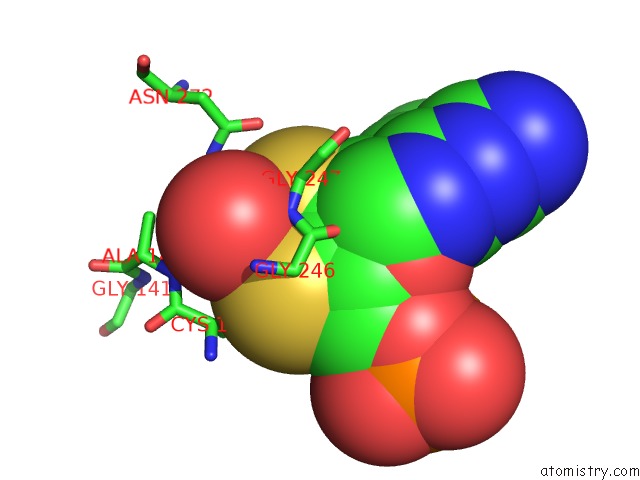
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 2 of Crystal Structure of Nitrate-Reducing Fragment of Assimilatory Nitrate Reductase From Pichia Angusta within 5.0Å range:
|
Reference:
K.Fischer,
G.Barbier,
H.-J.Hecht,
R.R.Mendel,
W.H.Campbell,
G.Schwarz.
Structural Basis of Eukaryotic Nitrate Reduction: Crystal Structures of the Nitrate Reductase Active Site Plant Cell V. 17 1167 2005.
ISSN: ISSN 1040-4651
PubMed: 15772287
DOI: 10.1105/TPC.104.029694
Page generated: Sun Aug 17 03:05:21 2025
ISSN: ISSN 1040-4651
PubMed: 15772287
DOI: 10.1105/TPC.104.029694
Last articles
Na in 4U9GNa in 4U7X
Na in 4U99
Na in 4U8W
Na in 4U6I
Na in 4U6B
Na in 4U44
Na in 4TWS
Na in 4U3Z
Na in 4U3E