Molybdenum »
PDB 5nqd-6op3 »
6bbl »
Molybdenum in PDB 6bbl: Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene
Enzymatic activity of Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene
All present enzymatic activity of Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene:
1.18.6.1;
1.18.6.1;
Protein crystallography data
The structure of Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene, PDB code: 6bbl
was solved by
O.A.Zadvornyy,
S.M.Keable,
J.Vertemara,
B.J.Eilers,
D.Karamatullah,
A.J.Rasmussen,
L.De Gioia,
G.Zampella,
L.C.Seefeldt,
J.W.Peters,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 39.76 / 1.68 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 76.781, 128.251, 107.272, 90.00, 109.11, 90.00 |
| R / Rfree (%) | 15.1 / 18.5 |
Other elements in 6bbl:
The structure of Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene also contains other interesting chemical elements:
| Magnesium | (Mg) | 1 atom |
| Iron | (Fe) | 32 atoms |
Molybdenum Binding Sites:
The binding sites of Molybdenum atom in the Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene
(pdb code 6bbl). This binding sites where shown within
5.0 Angstroms radius around Molybdenum atom.
In total 2 binding sites of Molybdenum where determined in the Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene, PDB code: 6bbl:
Jump to Molybdenum binding site number: 1; 2;
In total 2 binding sites of Molybdenum where determined in the Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene, PDB code: 6bbl:
Jump to Molybdenum binding site number: 1; 2;
Molybdenum binding site 1 out of 2 in 6bbl
Go back to
Molybdenum binding site 1 out
of 2 in the Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene
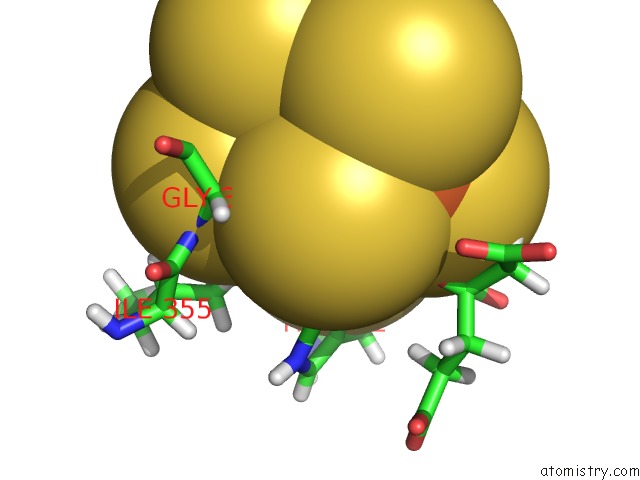
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 1 of Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene within 5.0Å range:
|
Molybdenum binding site 2 out of 2 in 6bbl
Go back to
Molybdenum binding site 2 out
of 2 in the Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene

Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Molybdenum with other atoms in the Mo binding
site number 2 of Crystal Structure of the A-96GLN Mofe Protein Variant in the Presence of the Substrate Acetylene within 5.0Å range:
|
Reference:
S.M.Keable,
J.Vertemara,
O.A.Zadvornyy,
B.J.Eilers,
K.Danyal,
A.J.Rasmussen,
L.De Gioia,
G.Zampella,
L.C.Seefeldt,
J.W.Peters.
Structural Characterization of the Nitrogenase Molybdenum-Iron Protein with the Substrate Acetylene Trapped Near the Active Site. J. Inorg. Biochem. V. 180 129 2017.
ISSN: ISSN 1873-3344
PubMed: 29275221
DOI: 10.1016/J.JINORGBIO.2017.12.008
Page generated: Sun Oct 6 16:29:51 2024
ISSN: ISSN 1873-3344
PubMed: 29275221
DOI: 10.1016/J.JINORGBIO.2017.12.008
Last articles
Ca in 3KMNCa in 3KM6
Ca in 3KMO
Ca in 3KM5
Ca in 3KLT
Ca in 3KL6
Ca in 3KLL
Ca in 3KHR
Ca in 3KLK
Ca in 3KHL